DNA Polymerase I
Importance of "proofreading" → DNA Polymerase I
|
|
DNA Polymerase → Proofreading steps:
1) Performs the first proofreading step just before a new nucleotide is covalently added to the growing chain.
|
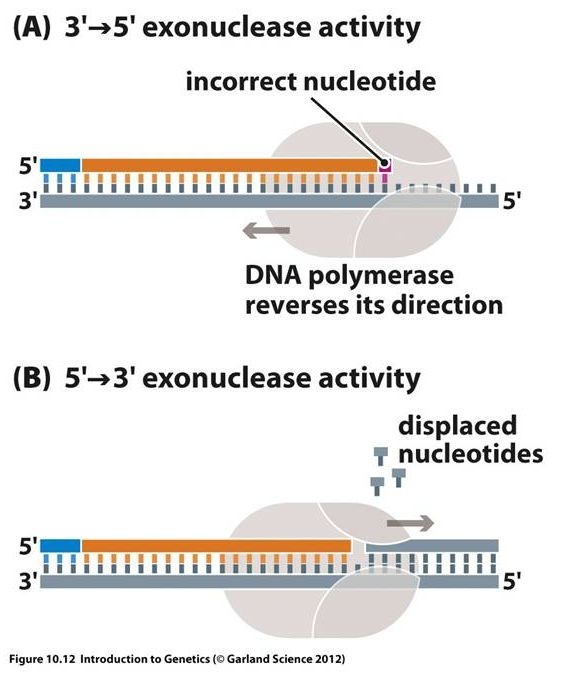
A mismatched base impedes translocation of DNA polymerase I to the next site. Sliding backwards the enzyme corrects the mistake with its 3'→5' exonuclease activity, then resume its polymerase activity in 5'→3' direction.
|
|
Klenow fragment of DNA polymerase I
|
Exonuclease activity enhances accuracy of DNA replication
3' to 5' exonuclease
3' to 5' exonuclease
- 3’-exonuclease activity removes nucleotides from 3’-end of growing chain – removes incorrect or mismatched bases
- 3’-exonuclease works slowly ca. polymerase activity
- Polymerase unable to extend mismatch pair – elongation stalls
- 3’ exonuclease therefore has time to catch up
- 3’-5’ Exonuclease activity acts as proof-reader
- 5’-3’ Exonuclease activity acts as editor
- 5’-exonuclease acts on duplex DNA, degrading it from 5’-end
- Can remove distorted segments in path of advancing polymerase
- Binds to nicks in dsDNA, moves in 5’ to 3’ direction, removing successive nucleotides
- 5’-exonuclease activity plays important role in DNA repair processes & in lagging strand synthesis
DNA Polymerise I → Major role not in DNA replication (not replicative polymerase)
- Until 1969, DNA polymerase I → only known DNA polymerase i E.coli, involved in DNA synthesis
- But, further characterisation → proved that enzyme's properties were not suitable for central role in replication!
- Maximal rate of DNA synthesis → (20 nucleotides/sec)
- At this rate, (460,000 sec/ 5.3 days) needed to replicate entire E.coli chromosome
- Too slow for an organism which divides every 20 mins!
- Around 400 molecules of DNA polymerase I per E.coli cell
- Excessive, since generally only 2 replication forks per cell
- Dissociates after catalysing the incorporation of 20-50 nucleotides
- However, shared this problem with every other known DNA polymerase (needs primer!)
DNA Polymerase Mutants (confirmed that Polymerase I is not the replicative polymerase!)
- DNA polymerase I coded for by the PolA1 gene
- De Luca and Cairns (1969) isolated E.coli mutants that lacked DNA polymerase I activity.
- Cells containing the polA1 mutation are viable/ more sensitive to UV light
- Mutation: codon for tryptophan 342 is mutated to a stop codon
- polA1 mutants grow normally - used to search for other DNA polymerase activities in E.coli (DNA polymerase II and III)
References
- Molecular Biology of the cell (6th ed)
- The molecules of life, Kuriyan et al
- Principles of Biochemistry, Lehninger (5th ed)
- Biochemistry, Berg et al (Internation ed)