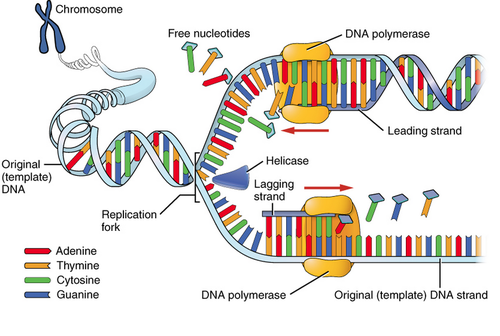
Leading and Lagging strands
|
|
|
|
The leading and Lagging strands are synthesised in a coordinated fashion...
- The duplex DNA is unwound by a hexameric helicase called DnaB.
- Copies of single-stranded-binding protein (SSB) binds to the unwound strands, keeping the strands separated so that both strands can serve as templates.
- Leading strand, a RNA primer (formed by primase) is only needed at the start of the replication, once replication fork established DNA polymerase is continuously presented with a base-pairing end on which to ass new nucleotides.
- DNA polymerase III begins synthesis of the leasing strand starting from RNA primer.
- Topoisomerase II concurrently introduces right-handed (negative) supercoils to avert a topological crisis.
- The synthesis of lagging strand (more complex) but coordinated with synthesis of leading strand. It is formed in pieces called (okazaki fragments)
- RNA primase lays down a RNA primer, then DNA polymerase III lays starts to add nucleotides starting from the primer, so each time DNA polymerase III completes a short DNA okazaki fragment, it must start synthesising a completely new fragment at a site further along the template strand, this mechanism depends on DNA primase which forms RNA primers (basic concept described in more detail below).
- DNA polymerase III haloenzyme interacts with the hexameric helicase DnaB (more about haloenzyme Pg 862 on Biochemistry, Berg et al, Internation ed).
- Lagging-strand template is looped out so that it passes through the polymerase site in one subunit of dimeric polymerase III in the same direction as that of leading-strand template in the other strand (5' to 3')
- DNA polymerase III lets go of the lagging-strand template after adding about 1000 nucleotides by releasing the sliding clamp.
- A new loop is then formed, a sliding clamp is added and primase again synthesises a short stretch of RNA primer to initate the formaton of another okazaki fragment.
- This mode of replication called trombone model, as size of loop lengthens and shorten like slide on a trombone.
- The gaps between fragments of lagging strand filled by DNA polymerase I. This enzyme uses 5' to 3' exonuclease activity to remove the RNA primer lying ahead of the polymerase site. (This can't be done by DNA polymerase II as it lacks 5' to 3' editing capability!)
- Finally DNA ligase connects the fragments.